 subis genome was relatively complete with 91% of avian orthologs detected as complete sequences (89.1% being single-copy and 1.9% being duplicated), which was in the range of other non-model avian genomes 18. The assembly length was similar to other avian genomes, which are typically between 1.0 and 1.2 Gb 17. The annotation included 12,686 genes (SI Appendix, Table S1). subis reference genome assembly based on long reads and linked reads was 1.17 Gb in length, consisted of 2896 scaffolds, had an N50 scaffold length of 6.13 Mb and an N50 contig length of 3.08 Mb. This study expands our understanding of the whole-genome contribution to migration and yields insight into migration timing behavior. Lastly, we conducted a genomic differentiation analyses between different migratory phenotypes to identify regions in the genome that may influence this trait. We also calculated polygenic scores (PGS) to assess the genetic predisposition of migratory timing. We examined results from genome-wide association studies (GWAS), to calculate the proportion of phenotype variation explained by our genomic data (PVE) along with presence of any associated single nucleotide polymorphisms (SNPs). Our objective was to examine the genomic architecture of migration timing by assembling a reference genome for the purple martin and integrating sequencing data with light-level geolocator tracks. For example, individuals breeding in the southern edge of the range in Florida may arrive as early as mid-January, while their northern counterparts in Alberta may arrive as late as June 16. 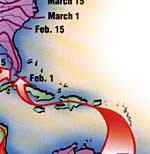
It is thus a powerful system to study migration genomics. The purple martin is a Nearctic-neotropical migrant that travels over 7000 km between North America and South America 15 and exhibits extensive latitudinal variation in migration timing. 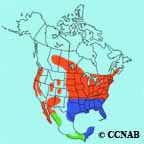
We overcame these limitations here, combining high resolution genomic data with an extensive migration tracking dataset for purple martins ( Progne subis subis), while specifically focusing on spring arrival timing given the importance of matching timing with resources critical for breeding. Another limitation associated with earlier studies was an inability to quantify migratory behavior in the wild-prior to 2007, it was not possible to track animals < 100 g on migration 14 and most migratory avian species fall into this size class. Previous genetic studies provide important insights regarding migration timing, such as in genes associated with circadian and circannual rhythms 7, 8, 9, 10, 11, 12 however, results vary across species 13 and are limited to small portions of the genome. However, migration timing is thought to be largely endogenously controlled 6, and thus knowledge of its genetic architecture (e.g., the identity, number, and location of genetic loci involved) is essential for predicting how migrants will respond to phenological changes that accompany climate change. Migration phenology may be influenced by many factors, including extrinsic factors such as rainfall 4 and wintering habitat conditions 5. These resources are becoming available earlier and it is unclear if migrants will be able to match these advances in timing, potentially leading to substantial population declines 3. For example, migrants must synchronize arrival at breeding grounds to coincide with seasonal resources availability 2. 
Overall, these results advance our understanding of the genomic underpinnings of migration timing.Ĭlimate change affects spring phenology in temperate zones and could have significant, negative impacts on migratory animals 1. On chromosome 1, a region that was differentiated between migration timing phenotypes contained genes that could facilitate nocturnal flights and act as epigenetic modifiers. A moderate to large amount of variance in spring migration arrival timing was explained by genomics (proportion of phenotypic variation explained by genomics = 0.74 polygenic score R 2 = 0.24). We examined the genetic architecture of migration timing in a long-distance migratory songbird (purple martin, Progne subis subis) by integrating genomic data with an extensive dataset of direct migratory tracks. Despite recent advances in our understanding of the underlying genetic basis of migration timing, the ways that migration timing phenotypes in wild individuals may map to specific genomic regions requires further investigation. The impact of climate change on spring phenology poses risks to migratory birds, as migration timing is controlled predominantly by endogenous mechanisms. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
